# All imports go here. Run me first!
import datetime
from pathlib import Path # Checks for files and so on
import numpy as np # Numpy for arrays and so on
import pandas as pd
import sys
import matplotlib.pyplot as plt # Matplotlib for plotting
# Ensure the plots are shown in the notebook
%matplotlib inline
import gdal
import osr
import numpy as np
%matplotlib inline
Fitting non-linear models¶
In the previous session, we looked at fitting linear models to observations. While this is a very common task, complex processes might require models which are non-linear.
One non-linear model is modelling LAI as a function of time (or temperature). In the Nothern Hemisphere, and for temperate latitudes, there is a clear seasonal cycle in vegetation, particularly visible in leaf area index (LAI). LAI dynamics can possibly be depicted by a “double logistic” curve. Mathematically, the double logistic looks like this
A double logistic
Mathematically, the function predicts the e.g. LAI (or some vegetation index) as
If we inspect this form, we can probably guess that \(p_0\) and \(p_1\) scale the vertical span of the function, whereas \(p_3\) and \(p_5\) are some sort of temporal shift, and the remaining parameters \(p_2\) and \(p_4\) control the slope of the two flanks. Something that will give rise to a self-respecting LAI curve might be
- \(p_0= 0.1\)
- \(p_1= 2.5\)
- \(p_2=0.19\)
- \(p_3= 120\)
- \(p_4= 0.13\)
- \(p_5= 220\)
Write a function that produces the double logistic when passed an array of time steps (e.g. 1 to 365), and an array with six parameters.
Do some plots and try to get some intuition on the model parameters!
A synthetic experiment¶
A first step is to do a synthetic experiment. This has the marked advantage of being a situation where we’re in control of everything.
t = np.arange(1, 366)
p = np.array([0.1, 2.5, 0.19, 120, 0.13, 220])
y = dbl_sigmoid_function(p, t)
yn = y + np.random.randn(len(t))*0.6
selector = np.random.rand(365)
passer = np.where(selector > 0.9, True, False)
tn = t[passer]
yn = yn[passer]
fig = plt.figure(figsize=(15, 4))
_ = plt.plot(t, y, '-', label="Ground truth")
_ = plt.plot(tn, yn, 'o', label="Simulated observations")
plt.legend(loc="best")
plt.xlabel("DoY")
plt.ylabel("LAI")
Text(0,0.5,'LAI')
/home/ucfajlg/miniconda3/envs/python3/lib/python3.6/site-packages/matplotlib/font_manager.py:1328: UserWarning: findfont: Font family ['sans-serif'] not found. Falling back to DejaVu Sans
(prop.get_family(), self.defaultFamily[fontext]))
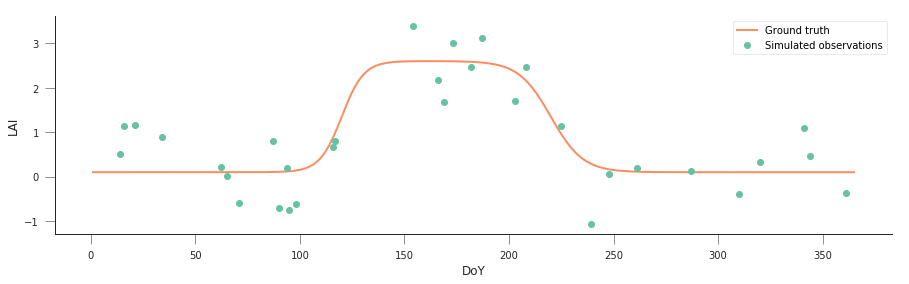
We know that the “true parameters” are given by
p = np.array([0.1, 2.5, 0.19, 120, 0.13, 220]), but we see that the
data is quite noisy and has significant gaps. As per last session, we
could try to modify the parameters “by hand”, and see how far we get,
but given that it’s 6, with different ranges, it looks a bit daunting.
Also, we’d need to assess how good the solution is for a particular set
of parameters, in other words, select a metric to quantify the goodness
of fit.
It is useful to consider a model of the incomplete, noisy observations of LAI (\(y_n\)) and the true value of LAI, \(y\). For overlapping time steps, the noisy data are just the “true” data plus some random Gaussian value with zero mean and a given variance \(\sigma_{obs}^2\) (in the experiment above, \(\sigma_{obs}=0.6\)):
Rearranging things, we have that \(y_n - y\) should be a zero mean Gaussian distribution with known variance. We have assumed that our model is \(f(\vec{p})=y\), so we can write the likelihood function, \(l(\vec{p})\)
It is convenient to take a logarithm of \(l(\vec{p})\), so that we have the log-likelihood:
<matplotlib.legend.Legend at 0x7fdd3698d358>
/home/ucfajlg/miniconda3/envs/python3/lib/python3.6/site-packages/matplotlib/font_manager.py:1328: UserWarning: findfont: Font family ['sans-serif'] not found. Falling back to DejaVu Sans
(prop.get_family(), self.defaultFamily[fontext]))
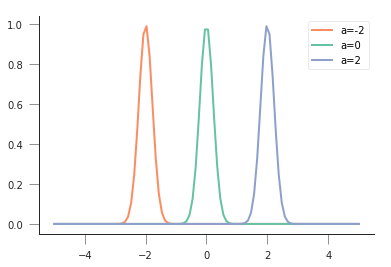
So for a sum of squares, the most likely result would be if all the mismatches were zero, which means that the log-likelihood is 0, and the likelihood, \(exp(0)=1\)!
However, the mismatch might not be 0, due to the added noise. So what we’re effectively looking for is a maximum in the log-likelihood, or a minimum of its negative as a function of \(\vec{p}\):
So, we can try our brute force guessing approach by minimising the cost function given by :math:`L(vec{p})`
The easiest way to obtain the solution is to use numerical optimisation
techniques to minimise the cost function. In scipy, there’s a good
selection of function
optimisers.
We’ll be looking at local optimisers: these will look for a minimum
in the vicinity of a user-given starting point, usually by looking at
the gradient of the cost function. The main function to consider here is
`minimise <https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html#scipy.optimize.minimize>`__.
Basically, minimize takes a cost function, a starting point, and
maybe extra arguments that are passed to the cost function, and uses one
of several algorithms to minimise the cost function. We import it with
from scipy.optimize import minimize
From the documentation,
minimize(fun, x0, args=(), method=None,
jac=None, hess=None, hessp=None,
bounds=None, constraints=(), tol=None,
callback=None, options=None)
Basically, fun is the name of the cost function. The first parameter
you pass to the cost function has to be an array with the parameters
that will be used to calculate the cost. x0 is the starting point.
args allows you to add extra parameters that are required for the
cost function (in our example, these would be
t, y_obs, passer, sigma_obs).
The minimize function returns an object with the
- Value of the function at the minimum,
- The value of the input parameters that attain the minimum,
- A message telling you whether the optimisation succeeded
- The number of iterations (
nit) and total function evaluations (nfev) - Some diagnostics
from scipy.optimize import minimize
from scipy.optimize import minimize
p0 = np.array([0, 5, 0.01, 90, 0.01, 200])
retval = minimize(cost_function, p0, args=(t, yn, passer, 0.6))
print(retval)
print ("********************************************")
if retval.success:
print("Optimisation successful!")
print(f"Value of the function at the minimum: {retval.fun:g}")
print(f"Value of the solution: {str(retval.x):s}")
fun: 21.342085853280015
hess_inv: array([[ 1.72923106e-02, -2.15349312e-02, 2.11636346e-03,
2.98572159e-02, 4.47526187e-03, -5.30376564e-02],
[-2.15349312e-02, 1.13986002e-01, -4.73509772e-02,
5.02075575e-01, -5.17466259e-02, -4.55944527e-01],
[ 2.11636346e-03, -4.73509772e-02, 7.51102753e-02,
-5.51690150e-01, 2.61989471e-02, 2.73170740e-01],
[ 2.98572159e-02, 5.02075575e-01, -5.51690150e-01,
9.54855368e+00, -3.19209970e-01, -3.30673084e+00],
[ 4.47526187e-03, -5.17466259e-02, 2.61989471e-02,
-3.19209970e-01, 5.74465509e-02, 3.44439220e-01],
[-5.30376564e-02, -4.55944527e-01, 2.73170740e-01,
-3.30673084e+00, 3.44439220e-01, 8.29858275e+00]])
jac: array([ 4.76837158e-07, 4.52995300e-06, -4.76837158e-07, -4.76837158e-07,
9.53674316e-07, 4.76837158e-07])
message: 'Optimization terminated successfully.'
nfev: 616
nit: 63
njev: 77
status: 0
success: True
x: array([2.15760787e-01, 2.22197385e+00, 2.73744085e-01, 1.23505589e+02,
2.63305511e-01, 2.22200329e+02])
****************************************
Optimisation successful
Value of the function at the minimum: 21.3421
Value of the solution: [2.15760787e-01 2.22197385e+00 2.73744085e-01 1.23505589e+02
2.63305511e-01 2.22200329e+02]
Do some synthetic experiments. For example:
Change the true parameters and see how the solution tracks the change.
Increase the added variance
Reduce or increase the number of observations
Use these experiments to challenge your understanding of the problem: Try to think what the expected result of these changes is, and write a set of functions that simplify the exploration.
Next: Real data¶
In the next session, you’ll be applying these techniques to MODIS LAI data